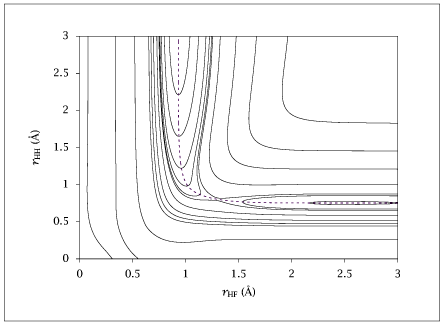
← Back to Figures Figure 5.3:
Contour representation of the potential-energy surface.
Code for Figure 5.3
Text of the GNU GPL .
initdata.py
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20 # Converted from initdata.m - returns parameter dict for LEPS potential
import numpy as np
def initdata ():
p = {}
p [ 'D1' ] = np . array ([ 129.3 , 94.8 , 129.3 ])
p [ 'B1' ] = np . array ([ 2.316 , 2.047 , 2.316 ])
p [ 'r10' ] = np . array ([ 0.931 , 0.753 , 0.931 ])
p [ 'D3' ] = np . array ([ 51.83 , 34.56 , 51.83 ])
p [ 'B3' ] = np . array ([ 2.229 , 1.522 , 2.229 ])
p [ 'r30' ] = np . array ([ 0.931 , 0.753 , 0.931 ])
p [ 'R_1' ] = np . array ([ 1.217 , 0.443 , 1.217 ])
p [ 'C' ] = np . array ([ 1174.9 , 388.8 , 1174.9 ])
p [ 'Si' ] = np . array ([ 3.090 , 1.929 , 3.090 ])
p [ 'A' ] = np . array ([ 1.315 , 0.762 , 1.315 ])
return p
if __name__ == '__main__' :
pass
inittrace.py
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60 # Converted from inittrace.m - compute reaction path trace for LEPS potential
# [depends] Vau.py rder.py lder.py
import numpy as np
from scipy.integrate import solve_ivp
import os
from Vau import Vau
from rder import rder
from lder import lder
def inittrace ( en1 , en2 , p ):
R1sar = 1.29781
R2sar = 0.80363
numr = 30
R1r = np . linspace ( R1sar , en1 , numr )
sqeps = np . sqrt ( np . finfo ( float ) . eps )
sol_r = solve_ivp ( lambda r1 , r2 : np . atleast_1d ( rder ( r1 , r2 [ 0 ], p )),
[ R1r [ 0 ], R1r [ - 1 ]], [ R2sar ],
method = 'BDF' , t_eval = R1r , rtol = sqeps , atol = sqeps )
R2r = sol_r . y [ 0 ]
R1sal = 1.297
R2sal = 0.80363
numl = 1000
R2l = np . linspace ( R2sal , en2 , numl )
sol_l = solve_ivp ( lambda r2 , r1 : np . atleast_1d ( lder ( r2 , r1 [ 0 ], p )),
[ R2l [ 0 ], R2l [ - 1 ]], [ R1sal ],
method = 'BDF' , t_eval = R2l , rtol = sqeps , atol = sqeps )
R1l = sol_l . y [ 0 ]
# reverse the 'r' branch
R1rn = R1r [:: - 1 ]
R2rn = R2r [:: - 1 ]
R1 = np . concatenate ([ R1rn , R1l ])
R2 = np . concatenate ([ R2rn , R2l ])
num = numl + numr
s = np . zeros ( num )
V = np . zeros ( num )
V [ 0 ] = Vau ( R1 [ 0 ], R2 [ 0 ], p )
for i in range ( 1 , num ):
s [ i ] = np . sqrt (( R1 [ i ] - R1 [ i - 1 ]) ** 2 + ( R2 [ i ] - R2 [ i - 1 ]) ** 2 ) + s [ i - 1 ]
V [ i ] = Vau ( R1 [ i ], R2 [ i ], p )
d3trace = np . zeros ( num * 3 )
for i in range ( num ):
j = i * 3
d3trace [ j ] = R1 [ i ]
d3trace [ j + 1 ] = R2 [ i ]
d3trace [ j + 2 ] = V [ i ]
return s , V , d3trace , num
if __name__ == '__main__' :
pass
Vau.py
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30 # Converted from Vau.m - London-Eyring-Polanyi-Sato (LEPS) potential
import numpy as np
def Vau ( r1 , r2 , p ):
r3 = r1 + r2
r = [ r1 , r2 , r3 ]
E1 = np . zeros ( 3 )
E3 = np . zeros ( 3 )
Q = np . zeros ( 3 )
J = np . zeros ( 3 )
for x in range ( 3 ):
E1 [ x ] = p [ 'D1' ][ x ] * ( np . exp ( - 2 * p [ 'B1' ][ x ] * ( r [ x ] - p [ 'r10' ][ x ])) -
2 * np . exp ( - p [ 'B1' ][ x ] * ( r [ x ] - p [ 'r10' ][ x ])))
E3a = p [ 'D3' ][ x ] * ( np . exp ( - 2 * p [ 'B3' ][ x ] * ( r [ x ] - p [ 'r30' ][ x ])) +
2 * np . exp ( - p [ 'B3' ][ x ] * ( r [ x ] - p [ 'r30' ][ x ])))
E3b = p [ 'C' ][ x ] * ( r [ x ] + p [ 'A' ][ x ]) * np . exp ( - p [ 'Si' ][ x ] * r [ x ])
E3 [ x ] = E3a if r [ x ] <= p [ 'R_1' ][ x ] else E3b
Q [ x ] = ( E1 [ x ] + E3 [ x ]) / 2
J [ x ] = ( E1 [ x ] - E3 [ x ]) / 2
retval = ( Q [ 0 ] + Q [ 1 ] + Q [ 2 ] -
np . sqrt ( 0.5 * (( J [ 0 ] - J [ 1 ]) ** 2 + ( J [ 1 ] - J [ 2 ]) ** 2 + ( J [ 0 ] - J [ 2 ]) ** 2 )))
return retval
if __name__ == '__main__' :
# Simple test - no file output for helper module
pass
rder.py
# Converted from rder.m - derivative dr2/dr1 along reaction path
# [depends] lder.py
import os
from lder import lder
def rder ( r1 , r2 , p ):
return 1.0 / lder ( r2 , r1 , p )
if __name__ == '__main__' :
pass
lder.py
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67 # Converted from lder.m - derivative dr1/dr2 along reaction path (London-Eyring-Polanyi-Sato)
import numpy as np
def lder ( r2 , r1 , p ):
r3 = r1 + r2
r = [ r1 , r2 , r3 ]
E1 = np . zeros ( 3 )
E3 = np . zeros ( 3 )
Q = np . zeros ( 3 )
J = np . zeros ( 3 )
dE1 = np . zeros (( 3 , 3 ))
dE3 = np . zeros (( 3 , 3 ))
for x in range ( 3 ):
E1 [ x ] = p [ 'D1' ][ x ] * ( np . exp ( - 2 * p [ 'B1' ][ x ] * ( r [ x ] - p [ 'r10' ][ x ])) -
2 * np . exp ( - p [ 'B1' ][ x ] * ( r [ x ] - p [ 'r10' ][ x ])))
E3a = p [ 'D3' ][ x ] * ( np . exp ( - 2 * p [ 'B3' ][ x ] * ( r [ x ] - p [ 'r30' ][ x ])) +
2 * np . exp ( - p [ 'B3' ][ x ] * ( r [ x ] - p [ 'r30' ][ x ])))
E3b = p [ 'C' ][ x ] * ( r [ x ] + p [ 'A' ][ x ]) * np . exp ( - p [ 'Si' ][ x ] * r [ x ])
E3 [ x ] = E3a if r [ x ] <= p [ 'R_1' ][ x ] else E3b
Q [ x ] = ( E1 [ x ] + E3 [ x ]) / 2
J [ x ] = ( E1 [ x ] - E3 [ x ]) / 2
# diagonal derivatives dE1[i,i] and dE3[i,i]
for i in range ( 3 ):
dE1 [ i , i ] = - 2 * p [ 'B1' ][ i ] * p [ 'D1' ][ i ] * (
np . exp ( - 2 * p [ 'B1' ][ i ] * ( r [ i ] - p [ 'r10' ][ i ])) -
np . exp ( - p [ 'B1' ][ i ] * ( r [ i ] - p [ 'r10' ][ i ])))
if r [ i ] <= p [ 'R_1' ][ i ]:
dE3 [ i , i ] = - 2 * p [ 'B3' ][ i ] * p [ 'D3' ][ i ] * (
np . exp ( - 2 * p [ 'B3' ][ i ] * ( r [ i ] - p [ 'r30' ][ i ])) +
np . exp ( - p [ 'B3' ][ i ] * ( r [ i ] - p [ 'r30' ][ i ])))
else :
dE3 [ i , i ] = p [ 'C' ][ i ] * np . exp ( - p [ 'Si' ][ i ] * r [ i ]) * (
1 - p [ 'Si' ][ i ] * ( r [ i ] + p [ 'A' ][ i ]))
dE1 [ 0 , 1 ] = 0. ; dE3 [ 0 , 1 ] = 0.
dE1 [ 1 , 0 ] = 0. ; dE3 [ 1 , 0 ] = 0.
dE1 [ 2 , 0 ] = dE1 [ 2 , 2 ]; dE3 [ 2 , 0 ] = dE3 [ 2 , 2 ]
dE1 [ 2 , 1 ] = dE1 [ 2 , 2 ]; dE3 [ 2 , 1 ] = dE3 [ 2 , 2 ]
dE1 [ 0 , 2 ] = 1000. ; dE3 [ 0 , 2 ] = 1000.
dE1 [ 1 , 2 ] = 1000. ; dE3 [ 1 , 2 ] = 1000.
dE1 [ 2 , 2 ] = 1000. ; dE3 [ 2 , 2 ] = 1000.
dQ = ( dE1 + dE3 ) / 2
dJ = ( dE1 - dE3 ) / 2
sq = np . sqrt ( 0.5 * (( J [ 0 ] - J [ 1 ]) ** 2 + ( J [ 1 ] - J [ 2 ]) ** 2 + ( J [ 0 ] - J [ 2 ]) ** 2 ))
dV1 = ( dQ [ 0 , 0 ] + dQ [ 2 , 0 ] - 0.5 / sq * (
J [ 0 ] * ( 2 * dJ [ 0 , 0 ] - dJ [ 2 , 0 ]) +
J [ 1 ] * ( - dJ [ 0 , 0 ] - dJ [ 2 , 0 ]) +
J [ 2 ] * ( 2 * dJ [ 2 , 0 ] - dJ [ 0 , 0 ])))
dV2 = ( dQ [ 1 , 1 ] + dQ [ 2 , 1 ] - 0.5 / sq * (
J [ 0 ] * ( - dJ [ 1 , 1 ] - dJ [ 2 , 1 ]) +
J [ 1 ] * ( 2 * dJ [ 1 , 1 ] - dJ [ 2 , 1 ]) +
J [ 2 ] * ( 2 * dJ [ 2 , 1 ] - dJ [ 1 , 1 ])))
return dV1 / dV2
if __name__ == '__main__' :
pass
main.py
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53 # Converted from hfc.m - contour plot data for LEPS potential surface
# [depends] initdata.py inittrace.py Vau.py
import numpy as np
from matplotlib import contour as mpl_contour
import os
from initdata import initdata
from inittrace import inittrace
from Vau import Vau
p = initdata ()
an1 , en1 = 0. , 3.
an2 , en2 = 0. , 3.
n1 , n2 = 301 , 301
step1 = ( en1 - an1 ) / ( n1 - 1 )
step2 = ( en2 - an2 ) / ( n2 - 1 )
x_arr = np . linspace ( an1 , en1 , n1 )
y_arr = np . linspace ( an2 , en2 , n2 )
z = np . zeros (( n2 , n1 ))
for p1 in range ( n1 ):
r1 = p1 * step1 + an1
for p2 in range ( n2 ):
r2 = p2 * step2 + an2
z [ p2 , p1 ] = Vau ( r1 , r2 , p )
levels = sorted ([ - 120 , - 110 , - 100 , - 95 , - 92 , - 90.5 , - 80 , - 60 , - 40 , - 20 ,
200 , 1000 , 3000 ])
# Compute contours using matplotlib (same algorithm as Octave contourc)
import matplotlib
matplotlib . use ( 'Agg' )
import matplotlib.pyplot as plt
fig , ax = plt . subplots ()
cs = ax . contour ( x_arr , y_arr , z , levels = levels )
plt . close ( fig )
with open ( 'hfc.dat' , 'w' ) as fid :
fid . write ( '# [contours] \n ' )
for i , level in enumerate ( cs . levels ):
for seg in cs . allsegs [ i ]:
fid . write ( f '# contour level { level } \n ' )
for row in seg :
fid . write ( f ' { row [ 0 ] : f } { row [ 1 ] : f } \n ' )
fid . write ( ' \n ' )
s , V , d3trace , num = inittrace ( en1 , en2 , p )
fid . write ( ' \n # [min] \n ' )
for i in range ( num ):
j = i * 3
fid . write ( f ' { d3trace [ j ] : f } { d3trace [ j + 1 ] : f } { d3trace [ j + 2 ] : f } \n ' )