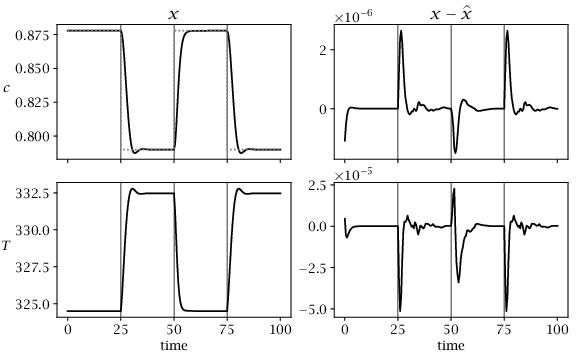
← Back to Figures Figure 4.7:
Closed-loop performance of combined nonlinear MHE/MPC with no disturbances. First column shows system states, and second column shows estimation error. Dashed line shows concentration setpoint. Vertical lines indicate times of setpoint changes.
Code for Figure 4.7
Text of the GNU GPL .
main.py
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221 # [makes] cstr_nonlinear_mpc_mhe.mat
#
# Control of a nonlinear CSTR using nonlinear MPC and MHE.
# Translated from cstr_nonlinear_mpc_mhe.m.
import numpy as np
import casadi
import scipy.io
import mpctools as mpc
def runcstr ( wmax , vmax ):
Delta = 0.5
Nx = 2 ; Nu = 1 ; Ny = 1 ; Nw = 1
Nsim = 200
Ncontroller = 10 ; Nmhe = 10
# CSTR parameters.
T0 = 350.0 ; c0 = 1.0 ; r_val = 0.219 ; k0 = 7.2e10 ; E = 8750.0
U = 54.94 ; rho = 1e3 ; Cp = 0.239 ; DeltaH = - 5e4
A = np . pi * r_val ** 2 ; rhoCp = rho * Cp ; h_val = 0.659 ; F = 0.1
def ode ( x , u ):
c , T , Tc = x [ 0 ], x [ 1 ], u [ 0 ]
k = k0 * np . exp ( - E / T )
rate = k * c
dcdt = F * ( c0 - c ) / ( A * h_val ) - rate
dTdt = ( F * ( T0 - T ) / ( A * h_val ) - DeltaH / rhoCp * rate
+ 2 * U / ( r_val * rhoCp ) * ( Tc - T ))
return np . array ([ dcdt , dTdt ])
cstrsim = mpc . getCasadiIntegrator ( ode , Delta , [ Nx , Nu ], [ 'x' , 'u' ],
funcname = 'cstrsim' , verbosity = 0 )
# Steady state (iterate a few steps to ensure dx/dt = 0).
xs = np . array ([ 0.878 , 324.5 ])
us = np . array ([ 300.0 ])
for _ in range ( 10 ):
xs = np . array ( cstrsim ( xs , us )) . flatten ()
C = np . array ([[ 0.0 , 1.0 ]]) # Measure temperature only.
G = np . array ([[ 1.0 ], [ 0.0 ]]) # Process noise enters concentration only.
ys = C @ xs # shape (1,)
CVs = [ 0 ] # Concentration is controlled variable (0-based).
# Bounds.
xlb = np . array ([ 0.1 , 250.0 ])
xub = np . array ([ 1.0 , 350.0 ])
umax_val = 0.25 * us [ 0 ]
ulb = us - umax_val
uub = us + umax_val
# CasADi functions.
print ( ' Building controllers.' )
flin = mpc . getCasadiFunc ( ode , [ Nx , Nu ], [ 'x' , 'u' ], funcname = 'flin' )
fmpc = mpc . getCasadiFunc ( ode , [ Nx , Nu ], [ 'x' , 'u' ], funcname = 'fmpc' ,
rk4 = True , Delta = Delta , M = 2 )
fsstarg = mpc . getCasadiFunc ( ode , [ Nx , Nu ], [ 'x' , 'u' ], funcname = 'fsstarg' )
x_sym = casadi . SX . sym ( 'x' , Nx )
u_sym = casadi . SX . sym ( 'u' , Nu )
w_sym = casadi . SX . sym ( 'w' , Nw )
fmhe = casadi . Function ( 'fmhe' , [ x_sym , u_sym , w_sym ],
[ fmpc ( x_sym , u_sym ) + casadi . mtimes ( G , w_sym )],
[ 'x' , 'u' , 'w' ], [ 'fmhe' ])
def measurement ( x ):
return x [ 1 ]
h_func = mpc . getCasadiFunc ( measurement , [ Nx ], [ 'x' ], funcname = 'h' )
# Linearize continuous model and compute LQR terminal cost matrix P.
ss = mpc . util . getLinearizedModel ( flin , [ xs , us ], names = [ 'A' , 'B' ], Delta = Delta )
A_lin , B_lin = ss [ 'A' ], ss [ 'B' ]
Q_mpc = 0.01 * np . diag ( xs **- 2 )
R_mpc = np . diag ( us **- 2 )
[ _ , P_dare ] = mpc . util . dlqr ( A_lin , B_lin , Q_mpc , R_mpc )
S_mpc = 10 * R_mpc
# MHE noise covariance inverses.
Qinv_mhe = 1.0 / 0.05
Rinv_mhe = 1.0 / 50.0
# Stage costs (closures capture Q_mpc, R_mpc, S_mpc, P_dare, Qinv, Rinv).
def l_ctrl ( x , u , Du , xsp , usp ):
dx = x - xsp ; du = u - usp
return ( mpc . mtimes ( dx . T , Q_mpc , dx ) + mpc . mtimes ( du . T , R_mpc , du )
+ mpc . mtimes ( Du . T , S_mpc , Du ))
l_ctrl_func = mpc . getCasadiFunc ( l_ctrl , [ Nx , Nu , Nu , Nx , Nu ],
[ 'x' , 'u' , 'Du' , 'x_sp' , 'u_sp' ],
funcname = 'l' )
def costtogo ( x , xsp ):
dx = x - xsp
return mpc . mtimes ( dx . T , P_dare , dx )
Pf_func = mpc . getCasadiFunc ( costtogo , [ Nx , Nx ], [ 'x' , 'x_sp' ], funcname = 'Pf' )
def lmhe_func ( w , v ):
return Qinv_mhe * w [ 0 ] ** 2 + Rinv_mhe * v [ 0 ] ** 2
lmhe = mpc . getCasadiFunc ( lmhe_func , [ Nw , Ny ], [ 'w' , 'v' ], funcname = 'l' )
# Assemble controller.
N_ctrl = { 'x' : Nx , 'u' : Nu , 't' : Ncontroller }
controller = mpc . nmpc ( f = fmpc , l = l_ctrl_func , N = N_ctrl , x0 = xs ,
Vf = Pf_func ,
par = { 'xsp' : xs , 'usp' : us },
lb = { 'x' : xlb , 'u' : ulb }, ub = { 'x' : xub , 'u' : uub },
guess = { 'x' : np . tile ( xs , ( Ncontroller + 1 , 1 )),
'u' : np . tile ( us , ( Ncontroller , 1 ))},
uprev = us ,
funcargs = { 'l' : [ 'x' , 'u' , 'Du' , 'x_sp' , 'u_sp' ]},
verbosity = 0 )
# Steady-state target finder.
N_ss = { 'x' : Nx , 'u' : Nu , 'y' : Ny }
sstarg_solver = mpc . sstarg ( fsstarg , h_func , N_ss ,
lb = { 'x' : xlb , 'u' : ulb }, ub = { 'x' : xub , 'u' : uub },
guess = { 'x' : xs , 'u' : us , 'y' : ys },
discretef = False , verbosity = 0 )
# Random disturbances (same seed each call to runcstr, matching Matlab behavior).
np . random . seed ( 0 )
urand = lambda n , m : 2 * np . random . rand ( n , m ) - 1
v = vmax * urand ( Ny , Nsim + 1 ) # (Ny, Nsim+1)
w_dist = wmax * urand ( Nw , Nsim ) # (Nw, Nsim)
y_init_perturb = vmax * urand ( Nmhe + 1 , Ny ) # (Nmhe+1, Ny)
# MHE.
y_init = np . tile ( ys , ( Nmhe + 1 , 1 )) + y_init_perturb # (Nmhe+1, Ny)
u_init = np . tile ( us , ( Nmhe , 1 )) # (Nmhe, Nu)
N_mhe = { 'x' : Nx , 'u' : Nu , 'w' : Nw , 'y' : Ny , 't' : Nmhe }
mhe = mpc . nmhe ( fmhe , h_func , u_init , y_init , lmhe , N_mhe ,
lb = { 'x' : xlb }, ub = { 'x' : xub },
guess = { 'x' : np . tile ( xs , ( Nmhe + 1 , 1 ))},
inferargs = True , verbosity = 0 )
# Closed-loop simulation.
x = np . full (( Nx , Nsim + 1 ), np . nan )
x [:, 0 ] = xs
y_meas = np . full (( Ny , Nsim + 1 ), np . nan )
u_ctrl = np . full (( Nu , Nsim ), np . nan )
xhat = np . full (( Nx , Nsim + 1 ), np . nan )
xtarg = np . full (( Nx , Nsim ), np . nan )
utarg = np . full (( Nu , Nsim ), np . nan )
# Setpoint: concentration steps down 10% every Nchange/2 steps.
t_idx = np . arange ( Nsim + 1 )
Nchange = 100
xsp = np . full (( Nx , Nsim + 1 ), np . nan )
xsp [ CVs [ 0 ], :] = xs [ CVs [ 0 ]] * ( 1 - 0.1 * ( np . mod ( t_idx , Nchange ) >= Nchange / 2 ))
xsp [:, - 1 ] = xsp [:, - 2 ]
u_prev = us . copy ()
for i in range ( Nsim + 1 ):
print ( '( %3d ) ' % ( i + 1 ), end = '' )
# Steady-state target.
sstarg_solver . fixvar ( 'x' , 0 , np . array ([ xsp [ CVs [ 0 ], i ]]), CVs )
sstarg_solver . solve ()
print ( 'Target: %s , ' % sstarg_solver . stats [ 'status' ], end = '' )
if sstarg_solver . stats [ 'status' ] != 'Solve_Succeeded' :
print ()
print ( 'Warning: sstarg failed at step %d !' % i )
break
if i < Nsim :
xtarg [:, i ] = np . array ( sstarg_solver . var [ 'x' , 0 ]) . flatten ()
utarg [:, i ] = np . array ( sstarg_solver . var [ 'u' , 0 ]) . flatten ()
sstarg_solver . saveguess ()
# Measurement and MHE.
y_meas [:, i ] = ( C @ x [:, i ]) + v [:, i ]
mhe . newmeasurement ( y_meas [:, i ], u_prev )
mhe . solve ()
print ( 'Estimator: %s , ' % mhe . stats [ 'status' ], end = '' )
if mhe . stats [ 'status' ] != 'Solve_Succeeded' :
print ()
print ( 'Warning: MHE failed at step %d !' % i )
break
xhat [:, i ] = np . array ( mhe . var [ 'x' , Nmhe ]) . flatten ()
mhe . saveguess ()
if i == Nsim :
print ( 'Done' )
break
# Controller.
controller . fixvar ( 'x' , 0 , xhat [:, i ])
controller . par [ 'x_sp' ] = [ xtarg [:, i ]] * ( Ncontroller + 1 )
controller . par [ 'u_sp' ] = [ utarg [:, i ]] * Ncontroller
controller . par [ 'u_prev' ] = [ u_prev ]
controller . solve ()
print ( 'Controller: %s ' % controller . stats [ 'status' ], end = '' )
if controller . stats [ 'status' ] != 'Solve_Succeeded' :
print ()
print ( 'Warning: controller failed at step %d !' % i )
break
u_ctrl [:, i ] = np . array ( controller . var [ 'u' , 0 ]) . flatten ()
controller . saveguess ()
u_prev = u_ctrl [:, i ] . copy ()
print ()
# Simulate plant.
x [:, i + 1 ] = ( np . array ( cstrsim ( x [:, i ], u_ctrl [:, i ])) . flatten ()
+ G [:, 0 ] * w_dist [ 0 , i ])
t = Delta * np . arange ( Nsim + 1 )
return dict ( x = x , u = u_ctrl , y = y_meas , xhat = xhat , xsp = xsp , t = t ,
v = v , w = w_dist , vmax = float ( vmax ), wmax = float ( wmax ))
wmaxes = [ 0 , 0.001 , 0.001 , 0.01 , 0.01 , 0.025 ]
vmaxes = [ 0 , 0.1 , 1 , 1 , 10 , 10 ]
data = []
for ( wmax_i , vmax_i ) in zip ( wmaxes , vmaxes ):
print ( 'Running wmax= %g , vmax= %g ' % ( wmax_i , vmax_i ))
data . append ( runcstr ( wmax_i , vmax_i ))
scipy . io . savemat ( 'cstr_nonlinear_mpc_mhe.mat' , { 'data' : data })